Apt-Get T7 Transcription Kit — 2'-Fluoro-Pyrimidine Set
T7 transcription kit for synthesis of nuclease-resistant RNAs, e.g. RNA aptamers.
Detailed Product Description
English Version
Deutsche Version
This kit contains a modified T7 RNA polymerase specifically engineered for efficient incorporation of 2'-fluoro modified nucleotides to RNA chains, as well as accessory reaction components, an optimized reaction buffer and a balanced NTP mix, where pyrimidines are 100% exchanged by their 2'-fluoro counterparts (2'F-CTP, 2'F-UTP). All purines (ATP, GTP) consist of 100% non-modified, high quality NTPs.
High quality and carefully balanced composition of all reaction components warrant synthesis of 2'F-pyrimidine modified RNA with high yield and efficiency. The kit is well suited for high yield synthesis of nuclease-resistant RNA and for RNA SELEX.
Introduction
Nucleic acids, in particular RNA, exhibit short half lifes, when exposed to nuclease-rich biological fluids, such as body and tissue fluids, animal sera etc. Chemical modifications of nucleotides and desoxyribonucleotides are a means to overcome the susceptibility of DNA and RNA to nucleolytic attack, and thus to increase nucleic acids stability in hostile environments.
Half life of RNA and DNA molecules in critical environments is greatly increased by replacing part of the incorporated nucleotides by chemically modified NTPs, such as 2'-fluoro NTPs. RNase A, the main causative endoribonuclease class for RNA-specific, nucleolytic activity, attacks RNA molecules at 3'-phosphates of pyrimidine nucleotides (C and U). 2'-fluoro modification of pyrimidines thus renders pyrimidine sites in RNA more resistant towards specific recognition and cleavage by RNase A.
Figure 1: Determining stability, enhanced nuclease resistance and prolonged half-life of 2'-pyrimidine (Py) RNA molecules (here: RNA aptamers) within nuclease-rich animal sera.
2'F-Py RNA aptamers (blue) and non-modified RNA aptamers (red) were incubated in nuclease-rich cell culture media containing fetal bovine serum. Radiolabelled full length RNA aptamers were separated by non-denaturing polyacrylamide gel electrophoresis (PAGE). Band intensities of full length aptamers were visualized and quantified by autoradiography.
Whereas, in this example, half-life of fully intact 2'F-Py modified RNA molecules is in the range of 60 min, the abundance of non-modified RNA molecules drops below detectable levels immediately upon introduction to the cell culture medium.
Generally, half-lifes of 2'F-Py RNA molecules (here: RNA aptamers) are strongly depended on individual sequence and secondary structure features, as well as on the length of the molecule, and thus may vary in a broad range. For certain 2'F-Py aptamers, half lifes were reported to exceed 24 hour time-frames. Data from Meyer C., Berg K. et al. (2014) RNA Biology 11 (1), 1–9.
Since wild-type enzymes, such as nucleases and polymerases, can not recognize modified NTPs as substrates, specifically engineered enzymes are required for incorporation of modified nucleotides into nucleic acid chains. Apt-Get 2'-F T7 RNA Polymerase incorporates 2'-fluoro modified nucleotides with high efficiency. Replacing all pyrimidines by their 2'-fluoro analogues, while retaining non-modified purines (ATP, GTP) offers an excellent balance between reaction efficiency, yield and nuclease resistance of synthetized, modified RNA.
Figure 2: T7 transcription with Apt-Get 2'-F T7 RNA Polymerase for incorporation of various 2'F-Py modified NTPs. In each lane, the denoted 2'-F NTP was used in spite of its non-modified counterpart. NON: Control reaction with all four standard NTPs, no modified NTPs are used. Transcript length: 500 nt RNA. 10 µl of each transcription assay was loaded on a 7 % [w/v] polyacrylamide gel with 8 M urea.
Modified nucleic acids, in particular RNA, containing 2'-fluoro pyrimidines exhibit pronounced stabilizing effects towards RNA secondary structure, due to increased impact of stacking forces. Secondary structure stability is improved, e.g. as compared to 2'-amino modified NTPs. However, safety issues might apply for in-vivo usage, since it cannot be excluded that fluoro-modified nucleotides get recycled into host DNA (Burmeister et al., Chem Biol. 2005 Jan;12(1):25-33).
High quality and carefully balanced composition of all reaction components warrant synthesis of 2'F-pyrimidine modified RNA with high yield and efficiency. The kit is well suited for high yield synthesis of nuclease-resistant RNA and for RNA SELEX.
Introduction
Nucleic acids, in particular RNA, exhibit short half lifes, when exposed to nuclease-rich biological fluids, such as body and tissue fluids, animal sera etc. Chemical modifications of nucleotides and desoxyribonucleotides are a means to overcome the susceptibility of DNA and RNA to nucleolytic attack, and thus to increase nucleic acids stability in hostile environments.
Half life of RNA and DNA molecules in critical environments is greatly increased by replacing part of the incorporated nucleotides by chemically modified NTPs, such as 2'-fluoro NTPs. RNase A, the main causative endoribonuclease class for RNA-specific, nucleolytic activity, attacks RNA molecules at 3'-phosphates of pyrimidine nucleotides (C and U). 2'-fluoro modification of pyrimidines thus renders pyrimidine sites in RNA more resistant towards specific recognition and cleavage by RNase A.
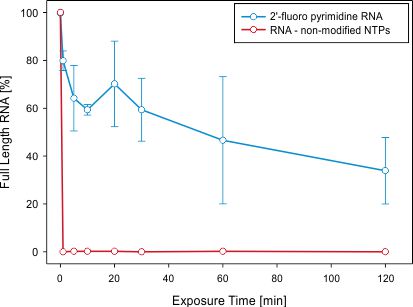
Figure 1: Determining stability, enhanced nuclease resistance and prolonged half-life of 2'-pyrimidine (Py) RNA molecules (here: RNA aptamers) within nuclease-rich animal sera.
2'F-Py RNA aptamers (blue) and non-modified RNA aptamers (red) were incubated in nuclease-rich cell culture media containing fetal bovine serum. Radiolabelled full length RNA aptamers were separated by non-denaturing polyacrylamide gel electrophoresis (PAGE). Band intensities of full length aptamers were visualized and quantified by autoradiography.
Whereas, in this example, half-life of fully intact 2'F-Py modified RNA molecules is in the range of 60 min, the abundance of non-modified RNA molecules drops below detectable levels immediately upon introduction to the cell culture medium.
Generally, half-lifes of 2'F-Py RNA molecules (here: RNA aptamers) are strongly depended on individual sequence and secondary structure features, as well as on the length of the molecule, and thus may vary in a broad range. For certain 2'F-Py aptamers, half lifes were reported to exceed 24 hour time-frames. Data from Meyer C., Berg K. et al. (2014) RNA Biology 11 (1), 1–9.
Since wild-type enzymes, such as nucleases and polymerases, can not recognize modified NTPs as substrates, specifically engineered enzymes are required for incorporation of modified nucleotides into nucleic acid chains. Apt-Get 2'-F T7 RNA Polymerase incorporates 2'-fluoro modified nucleotides with high efficiency. Replacing all pyrimidines by their 2'-fluoro analogues, while retaining non-modified purines (ATP, GTP) offers an excellent balance between reaction efficiency, yield and nuclease resistance of synthetized, modified RNA.
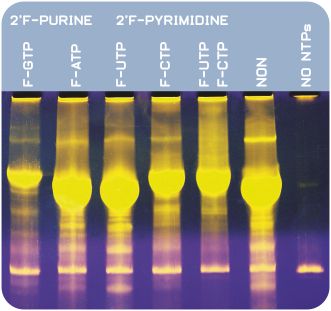
Figure 2: T7 transcription with Apt-Get 2'-F T7 RNA Polymerase for incorporation of various 2'F-Py modified NTPs. In each lane, the denoted 2'-F NTP was used in spite of its non-modified counterpart. NON: Control reaction with all four standard NTPs, no modified NTPs are used. Transcript length: 500 nt RNA. 10 µl of each transcription assay was loaded on a 7 % [w/v] polyacrylamide gel with 8 M urea.
Modified nucleic acids, in particular RNA, containing 2'-fluoro pyrimidines exhibit pronounced stabilizing effects towards RNA secondary structure, due to increased impact of stacking forces. Secondary structure stability is improved, e.g. as compared to 2'-amino modified NTPs. However, safety issues might apply for in-vivo usage, since it cannot be excluded that fluoro-modified nucleotides get recycled into host DNA (Burmeister et al., Chem Biol. 2005 Jan;12(1):25-33).

